
 |
 |
|||
Research
Cone Photoreceptor Biology and Age-related Macular Degeneration (AMD)
The success of future AMD therapies lies in their abilities to preserve, protect and eventually even restore cone photoreceptors, and their function, since it is ultimately the loss of these cells, and the sharp color vision they mediate, that results in devastating vision loss in AMD. The basis of such therapies relies on a thorough understanding of cone photoreceptors, the cells that support them [retinal pigment epithelial (RPE) cells] and their interactions within the human macula.
The macula is a unique and highly specialized region of the neural retina with high densities of rod and cone photoreceptor cells. These reach their highest densities at the center within the fovea, an approximately 2 mm diameter region. Macular degenerations are characterized by dysfunction and death of rod and cone photoreceptors and retinal pigment epithelial (RPE) cells in the macula. Studies suggest that perifoveal rod photoreceptor loss precedes that of cones in AMD; however, our understanding of the mechanism of macular and foveal cone loss in this process is poor. It appears that cones, especially the foveal cones, may have a propensity for survival in a region of the retina that has an overall predilection for degeneration. Our research program is focused on elucidating the precise role(s) of macular photoreceptors and retinal pigment epithelial cells in normal and diseased retinas.
Projects:
Fovea-associated gene discovery/expression profiling and candidate macular
dystrophy gene identification
To evaluate the role of macular and foveal cones in normal and diseased retina, and to gain insight into the biology of cone photoreceptors and the adjacent retinal pigment epithelium (RPE) that supports them, we have used multiple complimentary approaches for identifying genes enriched in the fovea from primate retinas. We developed a reference global expression profile or Transcriptome of the human posterior eye that evolved out of our need to query gene expression in the primate macula to support our studies of the pathobiology of AMD. This has yielded, EyeSAGE, which is an interactive tool and database for querying human retina and RPE gene expression. EyeSAGE simplifies the process of extracting useful information about genes expressed in the human retina and RPE from large gene expression datasets (Bowes Rickman et al. Invest. Ophthal. & Vis. Sci. 2006, in press) and is publicly accessible through the National Eye Institutes NEIBank EyeSAGE. EyeSAGE allows users to compare the pattern of gene expression in the retina and RPE to multiple tissues in order to identify genes that are likely to be expressed in a single cell type such as the cone photoreceptor. In addition, the database can be used for Genomic Convergence-combining expression data with available linkage information to generate lists of candidate genes for diseases such as macular degeneration.
Biochemical/functional studies of novel foveal-cone-associated proteins
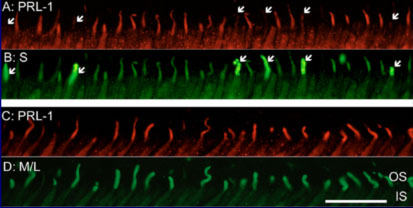
Studies of several of the proteins encoded by fovea-associated candidate genes are underway. One of these, encoded by a gene initially identified in our subtracted library, is a protein tyrosine phosphatase (PTP), PRL-1, specifically localized to the outer segments of red and green - but not blue - cone photoreceptors in the retina. PRL-1 is closely related to two other PTP's, PRL-2 and PRL-3 that are unique in the PTP superfamily in being C-terminal prenylated. Our studies implicate PRL-1 as a component of cellular response to photooxidative stress. Its oxidation, accompanied by the loss of its catalytic activity, and expression levels are induced in response to oxidative stress both in vitro and in vivo.
A mouse model for macular degeneration: Human APOE isoform knock-in Mice.
AMD is a late-onset, progressive, neurodegenerative disease with devastating impact on the elderly. This disease occurs primarily in people over the age of 65 years and accounts for approximately 50% of registered blindness in Western Europe and North America. AMD develops as either dry (atrophic) or wet (exudative). The pathogenesis of AMD is clearly multifactorial with genetic and environmental factors including aging, smoking, diet, gender, oxidative stress, and inflammation playing roles in onset and progression.
We developed a murine model of AMD by combining three of the risk factors for AMD: advanced age, apolipoprotein E isoform expression and exposure to a high-fat, high-cholesterol (HF-C) diet. The model recapitulates not only choroidal neovascularization (CNV) but also many of the RPE hallmarks of clinical AMD, which is unique in the field. Importantly, these changes require the presence of all three risk factors. This animal model of spontaneously-occurring CNV is also the first to incorporate physiologically-relevant risk factors of human disease. Using this model, both the mechanisms of pathology and therapeutic approaches for AMD can be studied.
 |
 |
Murine AMD model
Morphology of the RPE of AMD mouse (aged apoE4 TR mice maintained on a HF-C diet). Changes observed included RPE hyperpigmentation (A, arrow), RPE hypopigmentation (B, arrow), RPE atrophy (C, arrowheads), thick basal deposits. (D-G) Choroidal origin of neovascularization: Sequential 1 micron sections revealed CNV lesion confined between the RPE and BrM. (D) A capillary adjacent to a disruption in BrM (arrow). (E) Higher magnification of Inset in D showing apparent cellular migration at this break in BrM (arrow). (F) Double headed arrow shows the thinning of the ONL. (G) Higher magnification of Inset in F showing the splitting of BrM.
___________________________________________________________________________________
Home Research EyeSAGE Publications People Links