Estimation and Hypothesis Testing¶
In [1]:
library(tidyverse)
── Attaching packages ─────────────────────────────────────── tidyverse 1.2.1 ──
✔ ggplot2 2.2.1 ✔ purrr 0.2.5
✔ tibble 1.4.2 ✔ dplyr 0.7.5
✔ tidyr 0.8.1 ✔ stringr 1.3.1
✔ readr 1.1.1 ✔ forcats 0.3.0
── Conflicts ────────────────────────────────────────── tidyverse_conflicts() ──
✖ dplyr::filter() masks stats::filter()
✖ dplyr::lag() masks stats::lag()
In [2]:
options(repr.plot.width=4, repr.plot.height=3)
Set random number seed for reproucibility¶
In [3]:
set.seed(42)
Functions around probability distributions¶
Radnom numbers
In [4]:
rnorm(5)
- 1.37095844714667
- -0.564698171396089
- 0.363128411337339
- 0.63286260496104
- 0.404268323140999
In [5]:
x <- seq(-3, 3, length.out = 100)
plot(x, dnorm(x), type="l", ylab="PDF")

CDF
In [6]:
x <- seq(-3, 3, length.out = 100)
plot(x, pnorm(x), type="l", ylab="CDF")
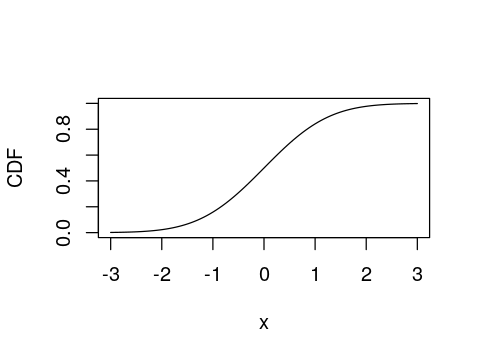
Quantiles (inverse CDF)
In [7]:
p = seq(0, 1, length.out = 101)
plot(p, qnorm(p), type="l", ylab="Quantile")

Point estimates¶
In [8]:
x <- rnorm(10)
In [9]:
x
- -0.106124516091484
- 1.51152199743894
- -0.0946590384130976
- 2.01842371387704
- -0.062714099052421
- 1.30486965422349
- 2.28664539270111
- -1.38886070111234
- -0.278788766817371
- -0.133321336393658
Mean¶
Manual calculation
In [10]:
sum(x)/length(x)
Using built-in function
In [11]:
mean(x)
Median¶
Manual calculation
In [12]:
x_sorted <- sort(x)
In [13]:
length(x)
Since there are an even number of observations, we need the average of the middle two data poitns
In [14]:
sum(x_sorted[5:6])/2
Using built-in function
In [15]:
median(x)
Quantiles¶
The mean is just the 50 percentile. We can use R to get any percentile we like.
In [16]:
quantile(x, 0.5)
In [17]:
quantile(x, seq(0,1,length.out = 5))
- 0%
- -1.38886070111234
- 25%
- -0.126522131318115
- 50%
- -0.0786865687327593
- 75%
- 1.45985891163508
- 100%
- 2.28664539270111
Exercise
Gene X is known to have a normal distribution with a mean of 100 units and a standard deviation of 15 units in the US population. With respect to this population,
- What is the medan value for gene X?
In [18]:
100
- What is the probability of finding a value of more than 130 for gene X if you pick a person at random?
In [19]:
m <- 100
s <- 15
In [20]:
1 - pnorm(130, m, s)
- If you measrue gene X and find that it is in the 95th percentile for this population, what is the measured value?
In [21]:
qnorm(0.95, m, s)
- Find answers to questions (1), (2) and (3) by simulating 1 million people sampled from the US population. Do they agree with the theoretical calculated values?
In [22]:
n <- 1e6
x <- rnorm(n, m, s)
In [23]:
median(x)
In [24]:
sum(x > 130)/n
In [25]:
quantile(x, 0.95)
- Plot the PDF of gene X using
ggplot2for values between 50 and 150. Give it a title ofPDF of N(100, 15), a subtile ofI made this!, and label the x-axis asGene Xand y-axis asPDF. Make the PDF blue, and fill the region under the curve blue with a transparency of 50%.
- Plot the PDF of gene X using
In [26]:
x <- 50:150
df <- data.frame(x=x, y=dnorm(x, m, s))
ggplot(df, aes(x=x, y=y)) +
geom_line(color='blue') +
geom_polygon(fill='blue', alpha=0.5) +
labs(title='PDF of N(100, 15)',
subtitle='I made this!',
x='Gene X',
y='PDF')

Interval estimates¶
Confidence intervals¶
In [27]:
ci = 0.95
In [28]:
alpha = (1-ci)
n <- length(x)
m <- mean(x)
s <- sd(x)
se <- s/sqrt(n)
me <- qt(1-alpha/2, df=n-1) * se
c(m - me, m + me)
- 94.215778757373
- 105.784221242627
Note that confidence intervals get larger as the confidence required increases.
In [29]:
ci = 0.99
In [30]:
alpha = (1-ci)
n <- length(x)
m <- mean(x)
s <- sd(x)
se <- s/sqrt(n)
me <- qt(1-alpha/2, df=n-1) * se
c(m - me, m + me)
- 92.3442793441823
- 107.655720655818
Making a function¶
Review of R custom functions¶
In [31]:
f <- function(a, b=1) {
a + b
}
In [32]:
f(2)
In [33]:
f(2,3)
In [34]:
f(b=4, a=1)
Exercise
Make a function called conf for calculating confidence intervals for
the sample mean that takes two arguments
- x is the vector of sample values
- ci is the confidence interval with a default of 0.95
The funciton should return a vector of two numbers indicating the lwoer and upper limeit of the confidence interval
Check that it gives the same answer as the example above.
In [35]:
conf <- function(x, ci=0.95) {
alpha = (1-ci)
n <- length(x)
m <- mean(x)
s <- sd(x)
se <- s/sqrt(n)
me <- qt(1-alpha/2, df=n-1) * se
c(m - me, m + me)
}
In [36]:
conf(x)
- 94.215778757373
- 105.784221242627
Coverage¶
In 1,000 experiments, we expect the true mean (0) to lie within the estimated 95% CIs 950 times.
In [37]:
n_expt <- 1000
n <- 10
cls <- t(replicate(n_expt, conf(rnorm(n))))
In [38]:
sum(cls[,1] < 0 & 0 < cls[,2])
Hypothesis testing¶
Binomial test¶
In [39]:
set.seed(123)
n = 50
tosses = sample(c('H', 'T'), n, replace=TRUE, prob=c(0.55, 0.45))
t = table(tosses)
t
tosses
H T
27 23
In [40]:
binom.test(table(tosses))
Exact binomial test
data: table(tosses)
number of successes = 27, number of trials = 50, p-value = 0.6718
alternative hypothesis: true probability of success is not equal to 0.5
95 percent confidence interval:
0.3932420 0.6818508
sample estimates:
probability of success
0.54
In [41]:
set.seed(123)
n = 250
tosses = sample(c('H', 'T'), n, replace=TRUE, prob=c(0.55, 0.45))
t = table(tosses)
t
tosses
H T
139 111
In [42]:
binom.test(t)
Exact binomial test
data: t
number of successes = 139, number of trials = 250, p-value = 0.0875
alternative hypothesis: true probability of success is not equal to 0.5
95 percent confidence interval:
0.4920569 0.6185995
sample estimates:
probability of success
0.556
What happens if we choose a one-sided test?¶
In [43]:
binom.test(t, alternative = "greater")
Exact binomial test
data: t
number of successes = 139, number of trials = 250, p-value = 0.04375
alternative hypothesis: true probability of success is greater than 0.5
95 percent confidence interval:
0.5020197 1.0000000
sample estimates:
probability of success
0.556
In [44]:
binom.test(t, alternative = "less")
Exact binomial test
data: t
number of successes = 139, number of trials = 250, p-value = 0.9668
alternative hypothesis: true probability of success is less than 0.5
95 percent confidence interval:
0.0000000 0.6089811
sample estimates:
probability of success
0.556
What happens if we change our null hypothesis?¶
In [45]:
binom.test(t, p = 0.55)
Exact binomial test
data: t
number of successes = 139, number of trials = 250, p-value = 0.8989
alternative hypothesis: true probability of success is not equal to 0.55
95 percent confidence interval:
0.4920569 0.6185995
sample estimates:
probability of success
0.556
Two-sample model¶
Welch t-test¶
In [46]:
set.seed(123)
n <- 10
x1 <- rnorm(n, 0, 1)
x2 <- rnorm(n, 1, 1)
In [47]:
t.test(x1, x2)
Welch Two Sample t-test
data: x1 and x2
t = -2.5438, df = 17.872, p-value = 0.02044
alternative hypothesis: true difference in means is not equal to 0
95 percent confidence interval:
-2.0710488 -0.1969438
sample estimates:
mean of x mean of y
0.07462564 1.20862196
Standard t-test¶
In [48]:
t.test(x1, x2, var.equal = TRUE)
Two Sample t-test
data: x1 and x2
t = -2.5438, df = 18, p-value = 0.02036
alternative hypothesis: true difference in means is not equal to 0
95 percent confidence interval:
-2.0705694 -0.1974232
sample estimates:
mean of x mean of y
0.07462564 1.20862196
Power of t-test¶
In [49]:
d <- 1 # Effect size is ratio of difference in means to standard deviation
library(pwr)
pwr.t.test(d = d, sig.level = 0.05, power = 0.9)
gives output
Two-sample t test power calculation
n = 22.02109
d = 1
sig.level = 0.05
power = 0.9
alternative = two.sided
NOTE: n is number in *each* group
Interpretation of the power calculaiton¶
If we did many experiments with n=23 per group where the effect size
is as specified and the test assumptions are valid, we expect that at
least 90% of them will have a p-value less than the nominal significance
level (0.05). If we used n=22 we would expect that just under 90% of
the experiments will have a p-value less than the nomial significance
level (0.05).
In particular notet that about 1-power of the experiments will fail to show a statistically significant p value even if the assumptionss are met (false negative).
In [50]:
n_expts <- 10000
n <- 22
alpha = 0.05
sum(replicate(n_expts, t.test(rnorm(n, 0, 1), rnorm(n, 1, 1))$p.value) < alpha)/n_expts
In [51]:
n_expts <- 10000
n <- 23
alpha = 0.05
sum(replicate(n_expts, t.test(rnorm(n, 0, 1), rnorm(n, 1, 1))$p.value) < alpha)/n_expts
Distribution of p-values under the null is uniform¶
That meas that you expect \(\alpha\) of the experiments to be false positives.
In [52]:
n_expt <- 10000
n <- 50
ps <- replicate(n_expts, t.test(rnorm(n), rnorm(n))$p.value)
In [53]:
sum(ps < alpha)/n_expts
In [54]:
hist(ps)

Paired and one-sample t-tests¶
A paired t-test is commonly used to evaluate if paired measuremetns (e..g. weight before and after a diet for the same person) has changed. The paired t-test is equivalent to a one-sample t-test that compares the difference in measurements for the paired values with a fixed number (usually 0).
In [55]:
x1 <- rnorm(10, 100, 15)
x2 <- rnorm(10, 100, 15)
delta <- x1 - x2
In [56]:
t.test(x1, x2, paired=TRUE)
Paired t-test
data: x1 and x2
t = -2.4618, df = 9, p-value = 0.03605
alternative hypothesis: true difference in means is not equal to 0
95 percent confidence interval:
-12.3446069 -0.5215923
sample estimates:
mean of the differences
-6.4331
In [57]:
t.test(delta, mu=0)
One Sample t-test
data: delta
t = -2.4618, df = 9, p-value = 0.03605
alternative hypothesis: true mean is not equal to 0
95 percent confidence interval:
-12.3446069 -0.5215923
sample estimates:
mean of x
-6.4331
Exercise
Suppose that the null hypotehsis is that there is no difference in the paired measurements and the standard deviation of the differene is 2.
- Run a simulation for 100,000 experiments with
n=25per experiment to show the distributon of the p values using a paired or one -sample t-test under the null. - If the significance level is 0.05, how many false positive results were observed?
In [58]:
n_expt <- 100000
n <- 25
s <- 2
ps <- replicate(n_expts, t.test(rnorm(10, 0, s), mu=0)$p.value)
In [59]:
hist(ps)

In [60]:
n <- 100000
s <- sqrt(2)
sd(rnorm(n, 0, s) - rnorm(n, 0, s))